library(tidyverse)
library(zoo)
library(gridExtra)
source(here::here("R/dives.R"))
depth <- read_csv(here::here("data/23A1302/out-Archive.csv"), show_col_types = FALSE) %>%
drop_na(Depth) %>%
mutate(Date = as.POSIXct(Time, format = "%H:%M:%S %d-%b-%Y", tz = "UTC")) %>%
rename(no_zoc_depth = Depth,
Depth = `Corrected Depth`)
dives <- get_dives(depth, 5, 80)Fine scale
First, read in the dive data and split into individual dives.
Next, load in the acoustic data. We have two kinds of acoustic data: manual annotations of sperm whale bouts, and triton-detected sperm whale clicks (aka, TPWS). We want to compare them and make sure they look similar.
to-do: update the manual annotations, they are newly validated.
source(here::here("R/broad_funcs.R"))
#sw_comb_bouts has just sperm whales, and all nearby bouts are combined.
manual <- read.csv(here::here("data/sw_comb_bouts.csv"))
#convert times to POSIXct and add boutID
manual <- manual %>%
mutate(start = ymd_hms(start, tz = "UTC"),
end = ymd_hms(end, tz = "UTC"),
bout = row_number())
window_hr <- 12
sw_manual <- tibble(date = seq(first(manual$start), last(manual$end), by = window_hr * 3600)) %>%
mutate(SW = map_dbl(date, \(d) sum_odon_exposure(data = manual, d)))
#read in each triton-found click with RL measurement (MPP, measured in dB)
tpws <- read.csv(here::here("data/sw/tpws.csv"), header = FALSE) %>%
rename(clicktime = V1, MPP = V2)
#convert the matlab dates to UTC
tpws <- tpws %>%
mutate(clicktime = as.POSIXct((clicktime - 719529) * 86400,
origin = "1970-01-01",
tz = "UTC"))
ggplot(tpws, aes(clicktime)) +
geom_histogram(color = "white", fill = "gray") +
geom_line(data = sw_manual, aes(date, SW * 800), color = "hotpink") +
theme_bw()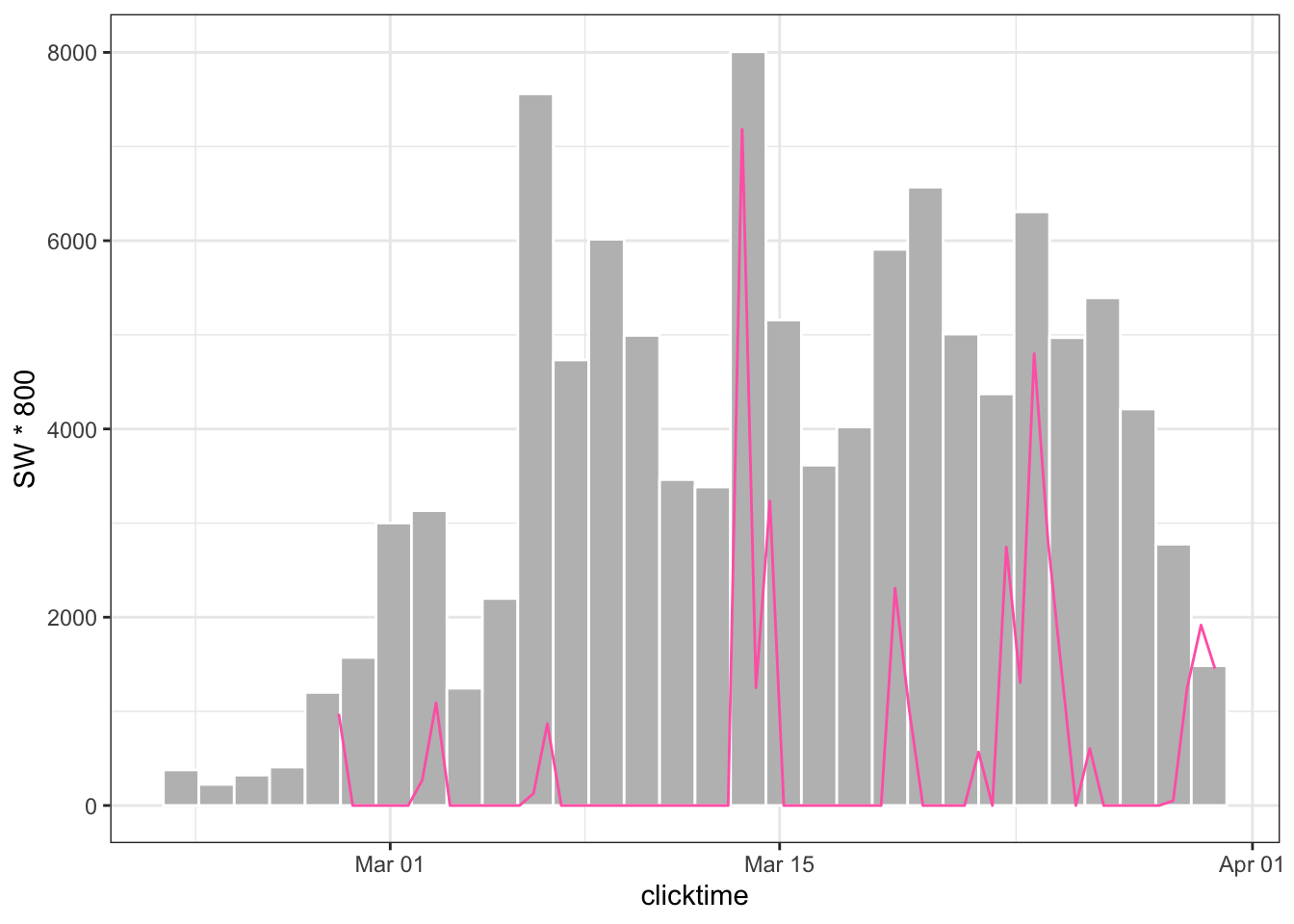
Cutting tpws to the manual annotations, but show 5 hours of tpws + dives on either side.
#filter tpws to only include times in the validated manual annotations
tpws_val <- manual %>%
mutate(data = map2(start - hours(5), end + hours(5), ~ {
tpws %>%
filter(clicktime >= .x & clicktime <= .y)
})) %>%
unnest(data) %>%
select(bout, start, end, clicktime, MPP)
# #choose window in minutes, then create columns for the window in the data frame
window_min <- 5
tpws_val <- tpws_val %>%
mutate(time_elapsed = difftime(clicktime, start, units = "secs"),
window = floor(as.numeric(time_elapsed / (window_min * 60))),
win_start = start + seconds(window * window_min * 60))
# #calculate the average received level for each window
avg_Rl <- tpws_val %>%
group_by(bout, win_start) %>%
summarize(mean_MPP = mean(MPP, na.rm = TRUE),
clicks_per_bin = n(),
.groups = "drop")
#put the window average RLs into the tpws data frame
tpws_full <- tpws_val %>%
inner_join(avg_Rl, by = "win_start")
#get the dive starts so that we can also calcualte mean RL for each dive
dive_starts <- dives %>%
group_by(dive_id) %>%
slice(1) %>%
ungroup() %>%
select(dive_id, Date)
#add a column to the twps dataframe which has the associated dive ID for each detection
tpws_full$dive_id <- findInterval(tpws_full$clicktime, dive_starts$Date)Calculating when the device was NOT recording audio.
source(here::here("R/non_recording.R"))
#wav_timestamps.csv is the output of the function get_wav_timestamps.R.
recording <- read.csv(here::here("data/wav_timestamps.csv")) %>%
mutate(start = ymd_hms(start_time, tz = "UTC"),
end = ymd_hms(end_time, tz = "UTC")) %>%
drop_na(end)
non_recording <- calculate_non_recording(recording)Plotting the received level vs. time on top, with the dive record underneath.
source(here::here("R/fine_plots.R"))
#make the plots
bout_plots <- map(1:nrow(manual),
~make_bout_plot(.x, tpws_full, depth, non_recording, manual))
bout_plots[[1]]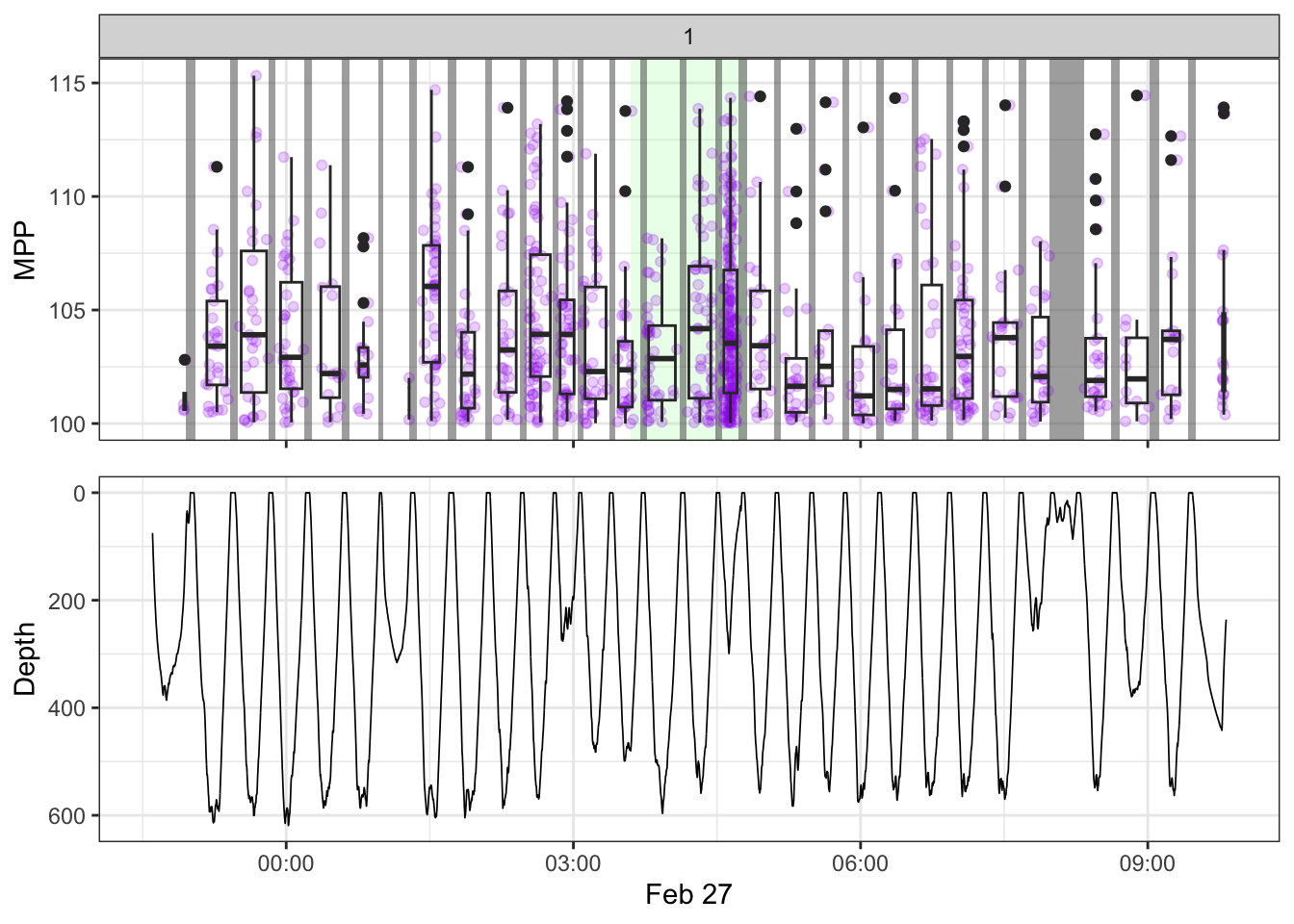
[[2]]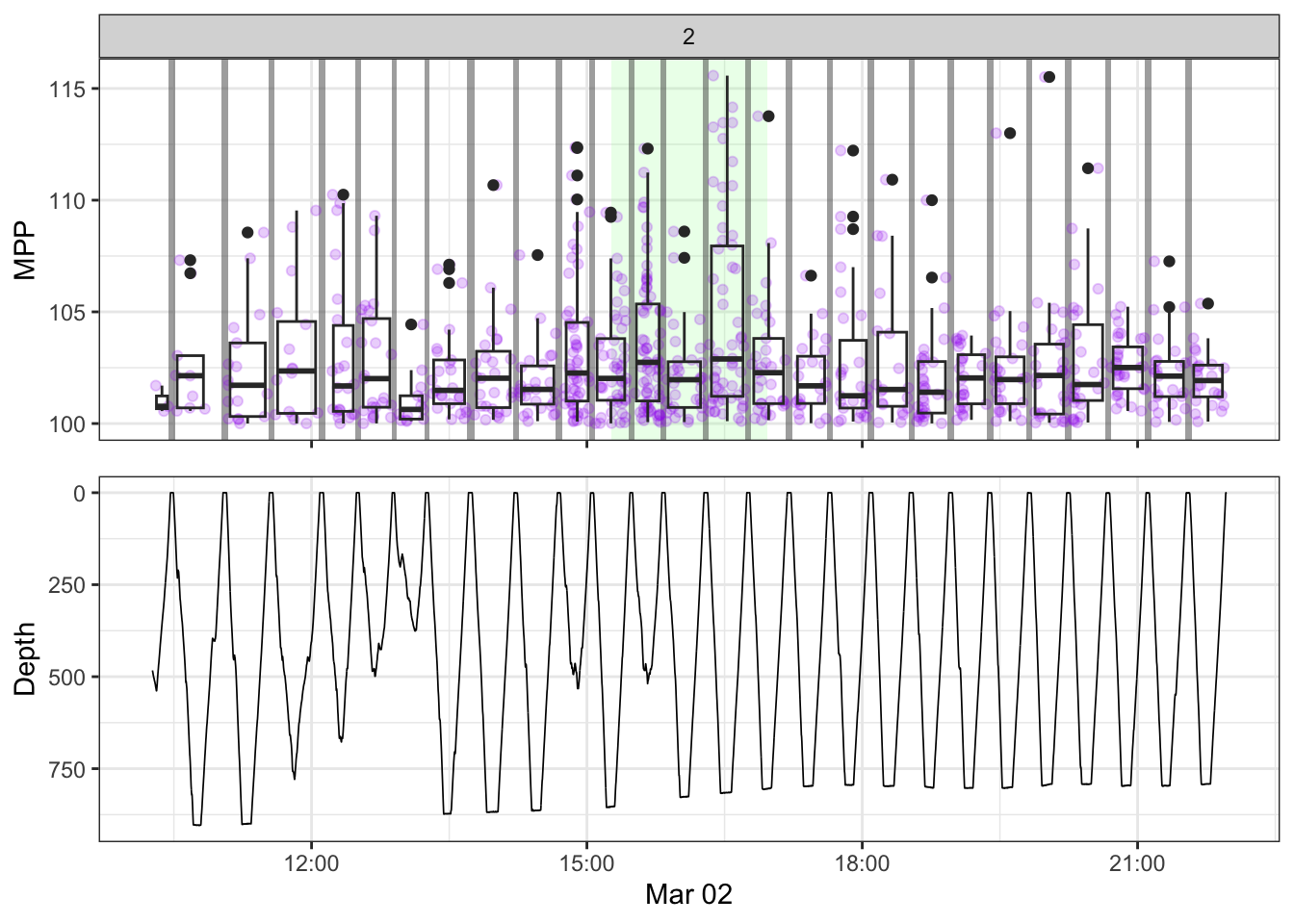
[[3]]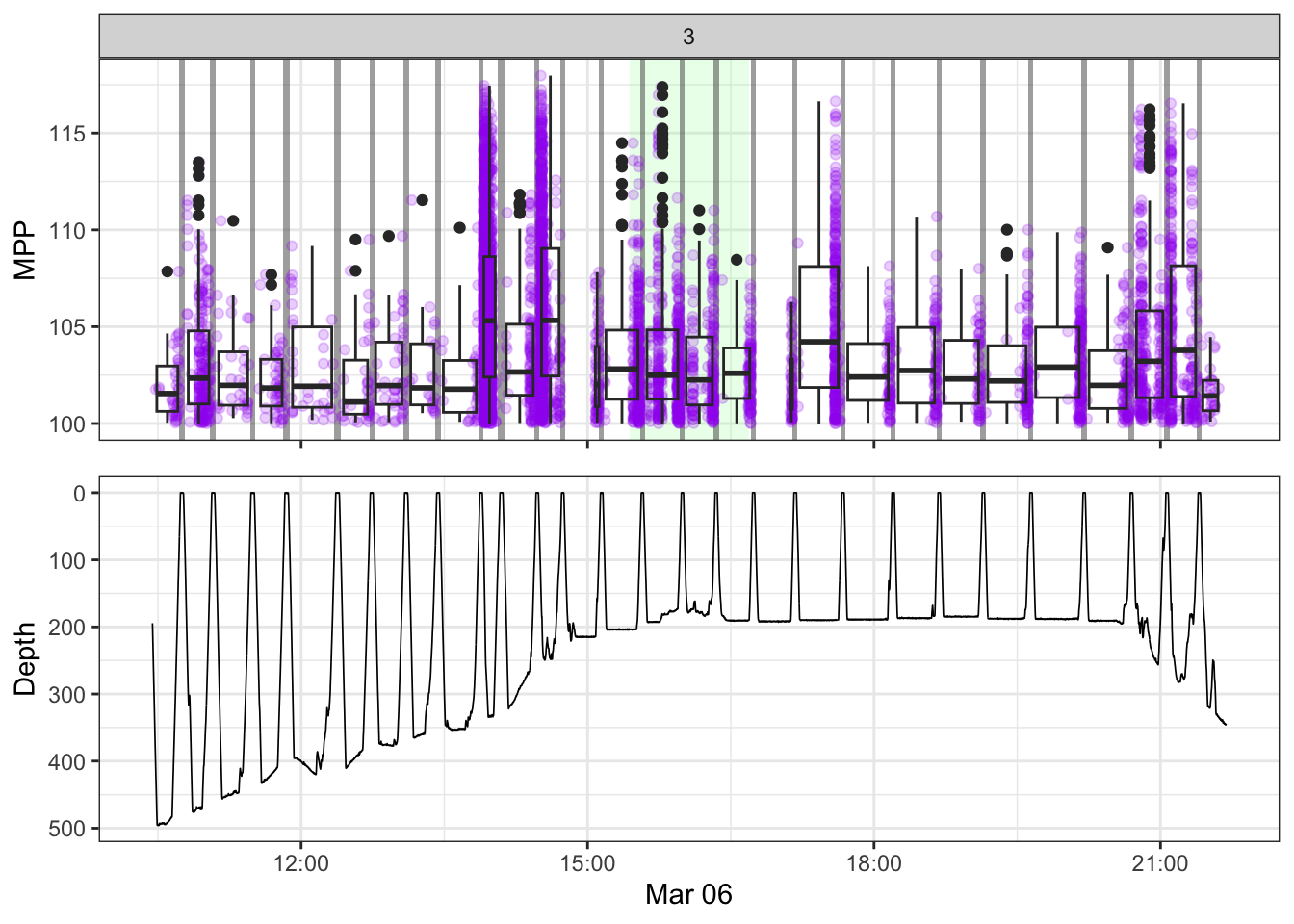
[[4]]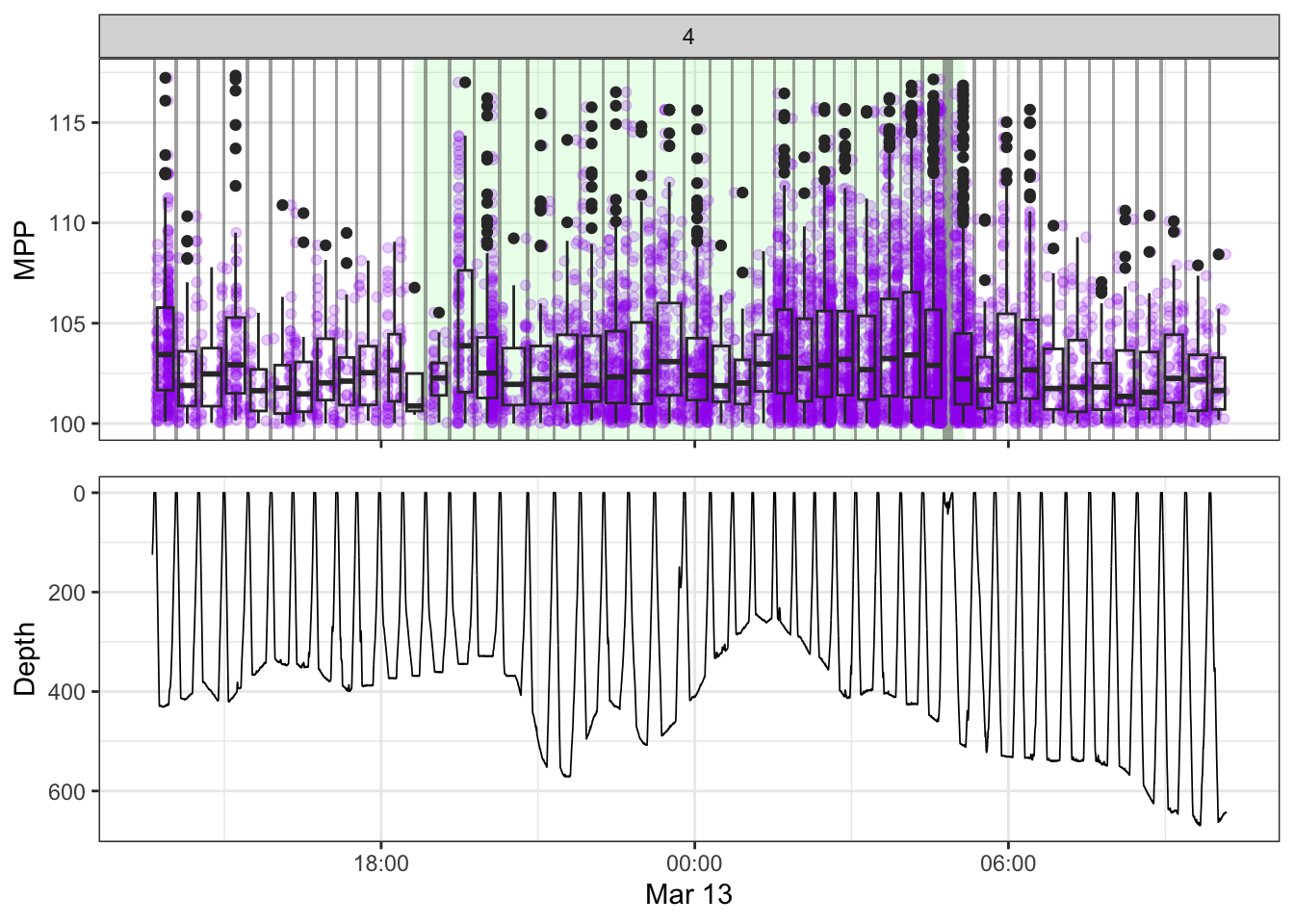
[[5]]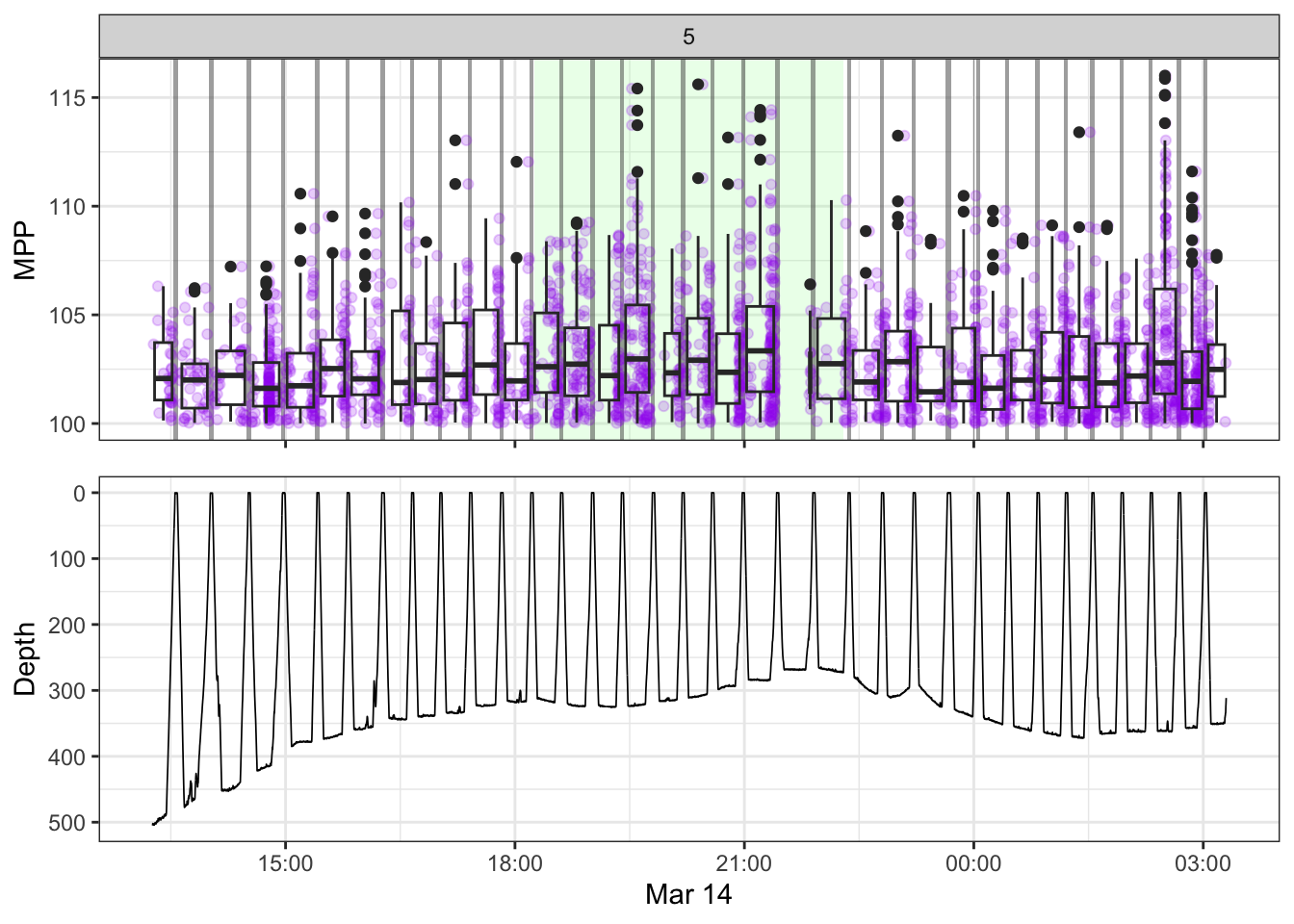
[[6]]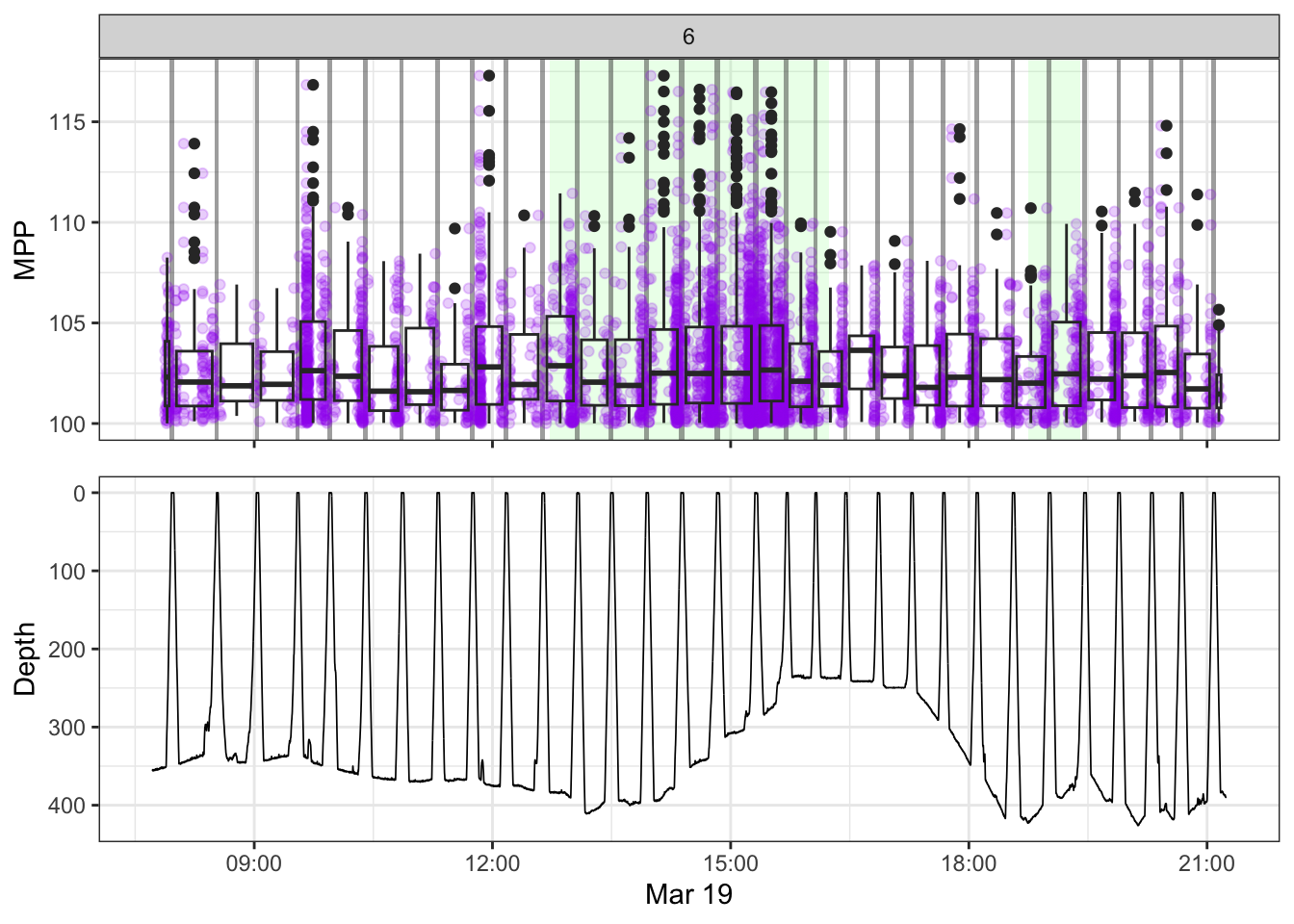
[[7]]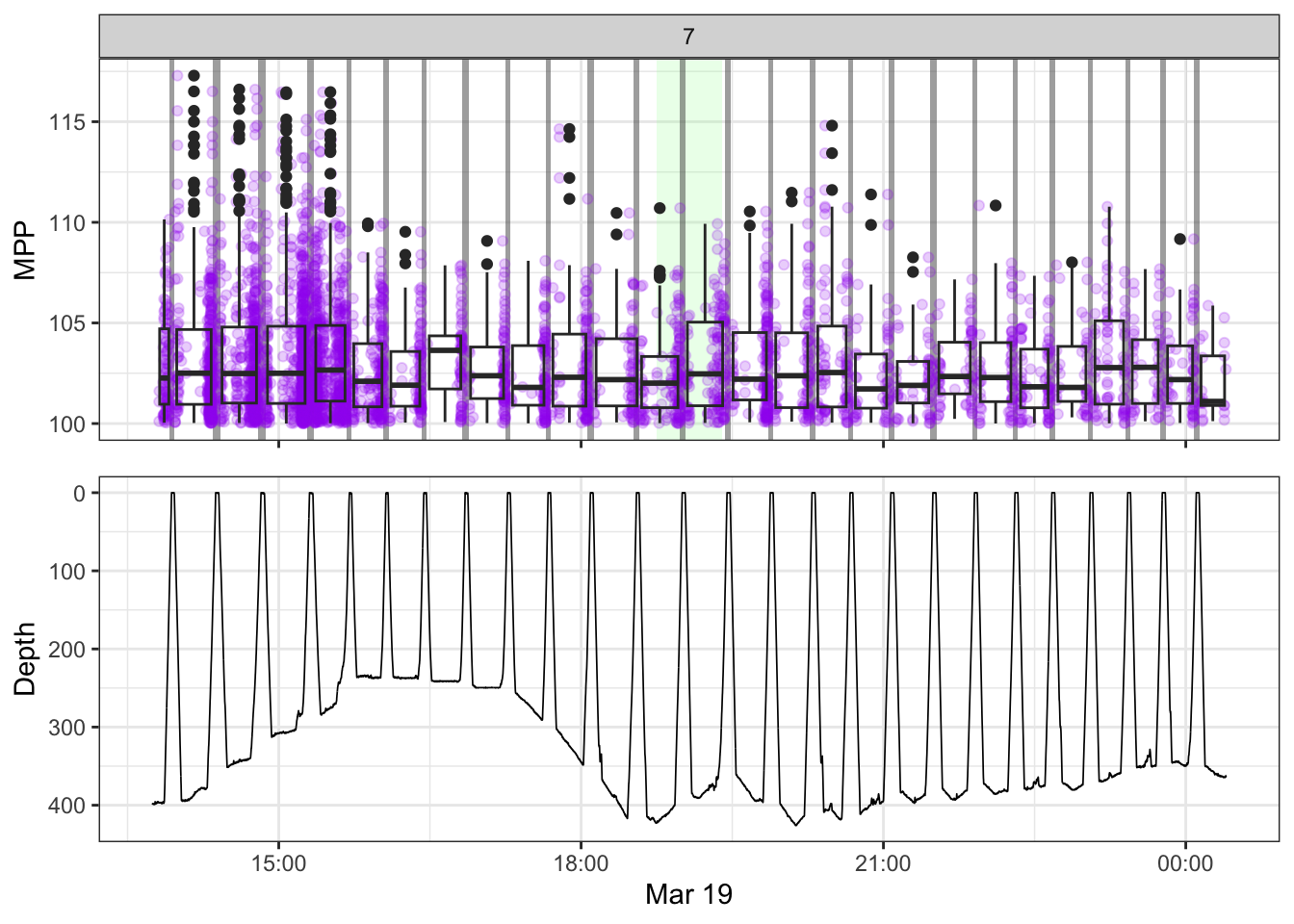
[[8]]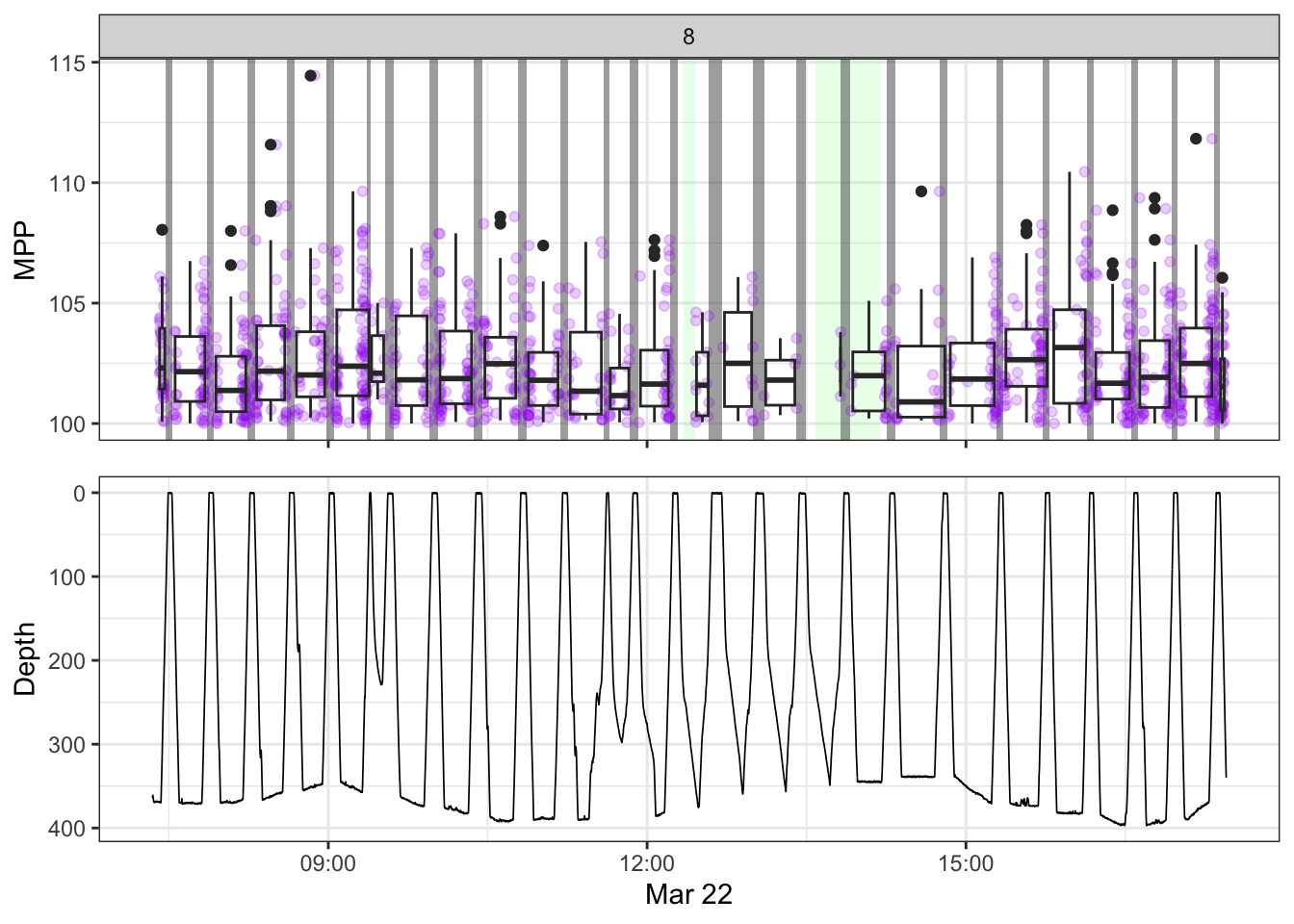
[[9]]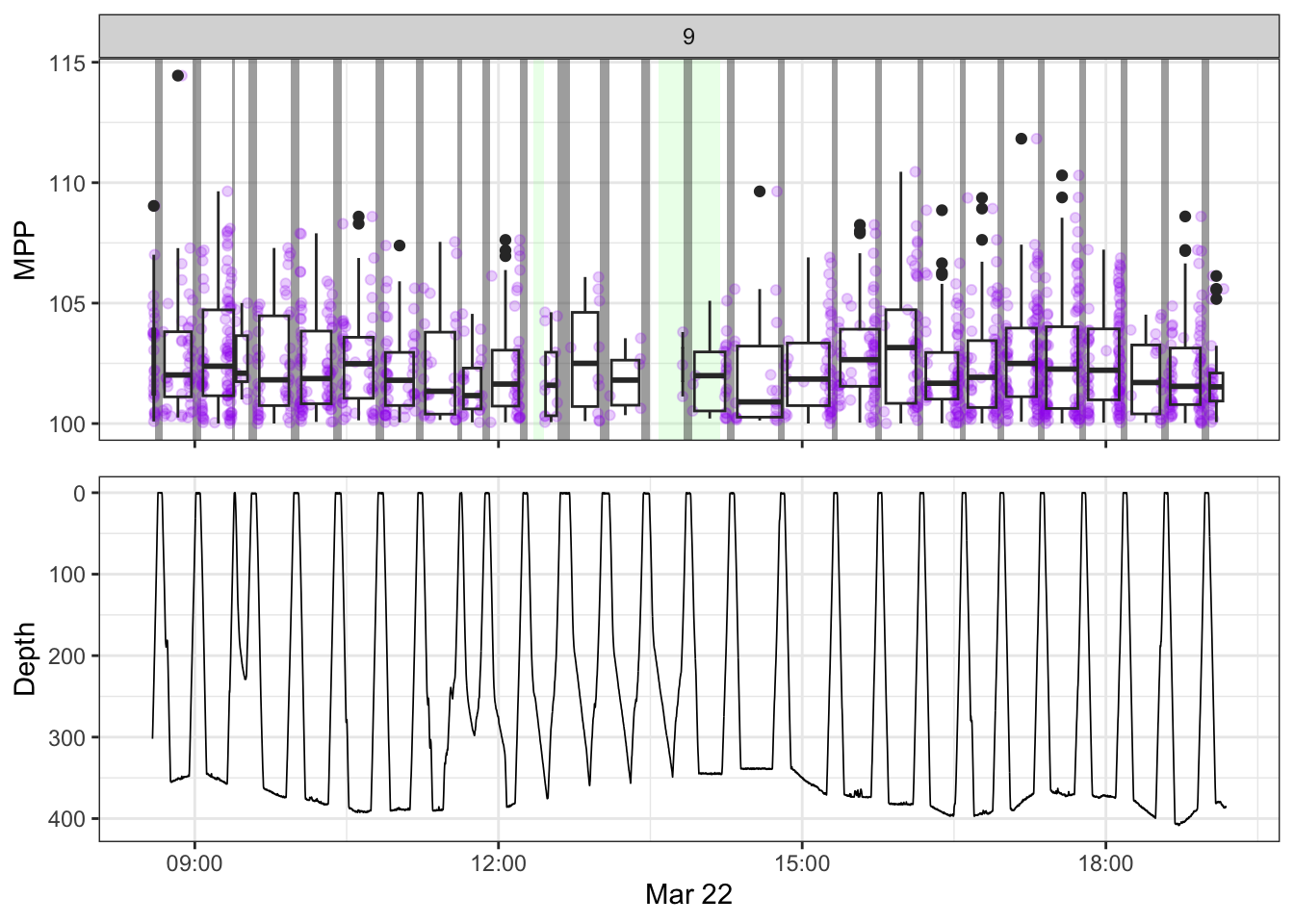
[[10]]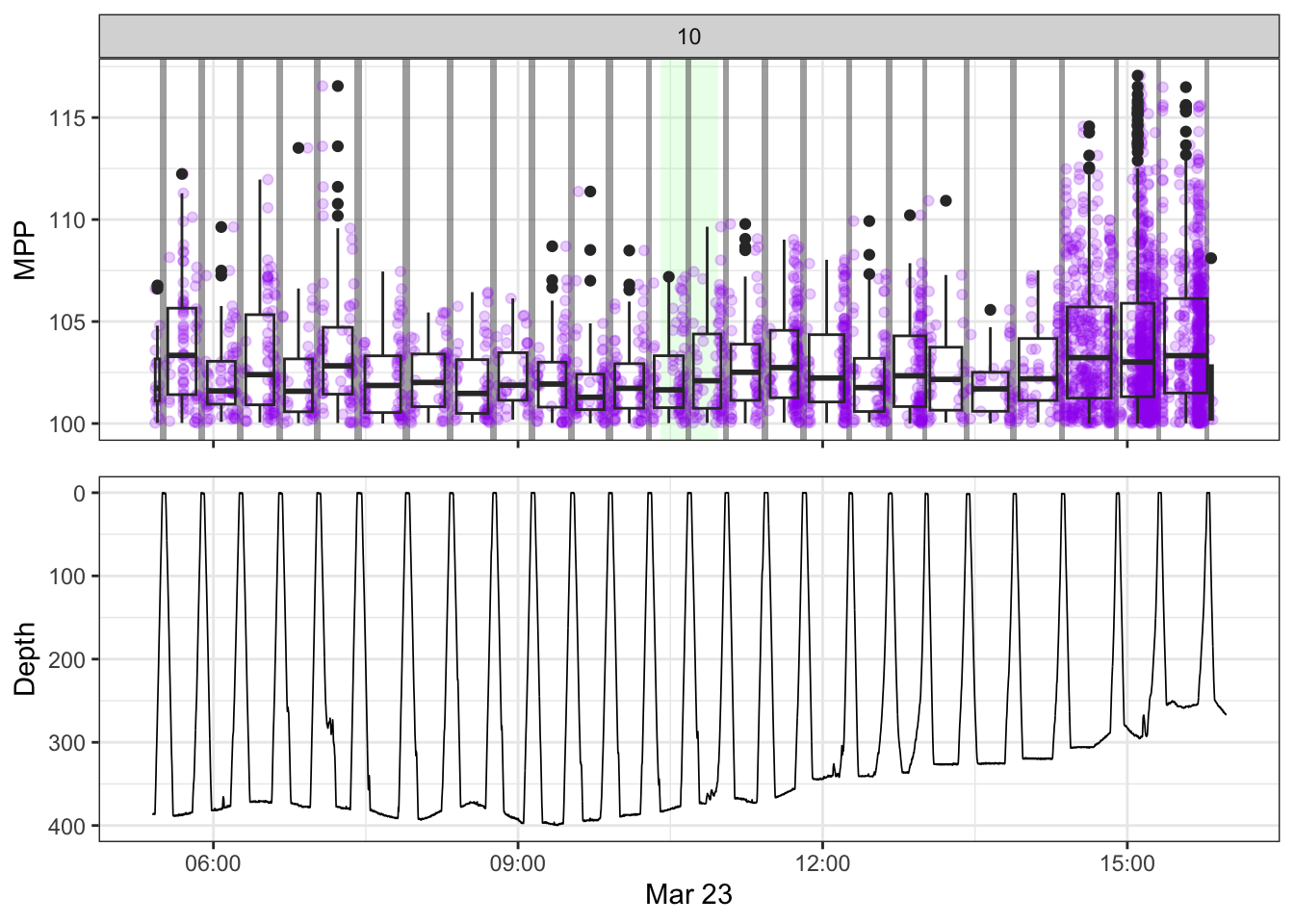
[[11]]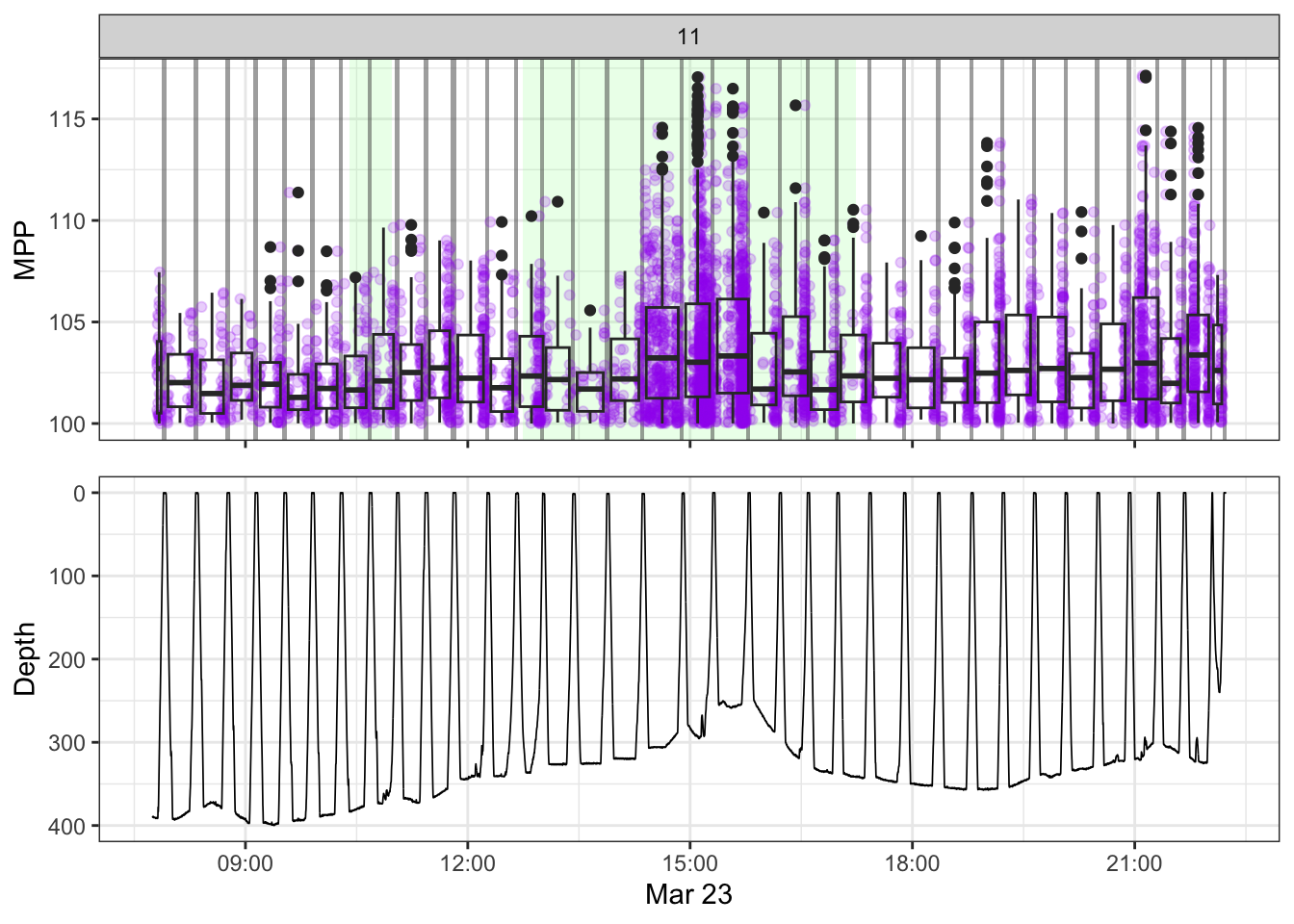
[[12]]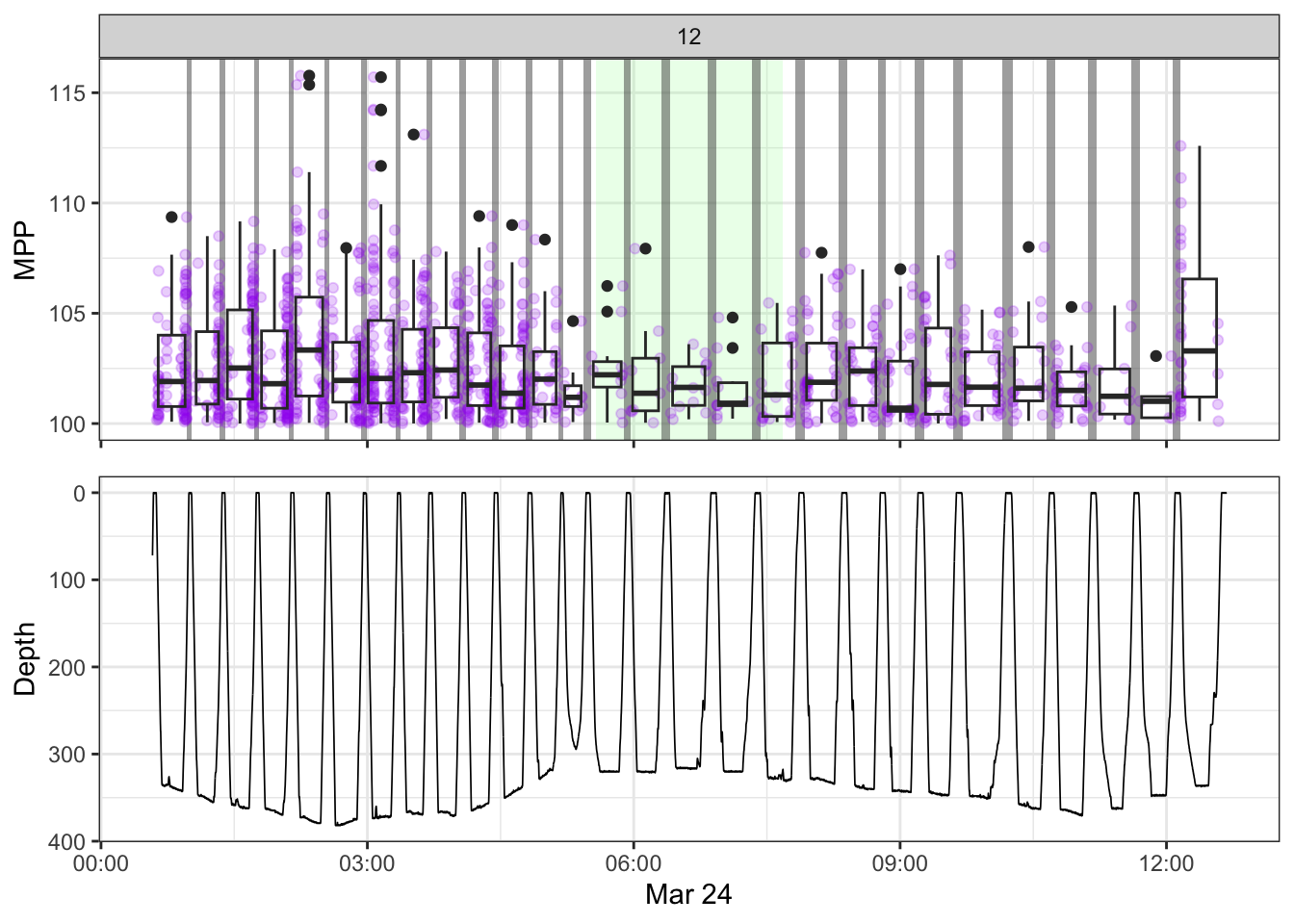
[[13]]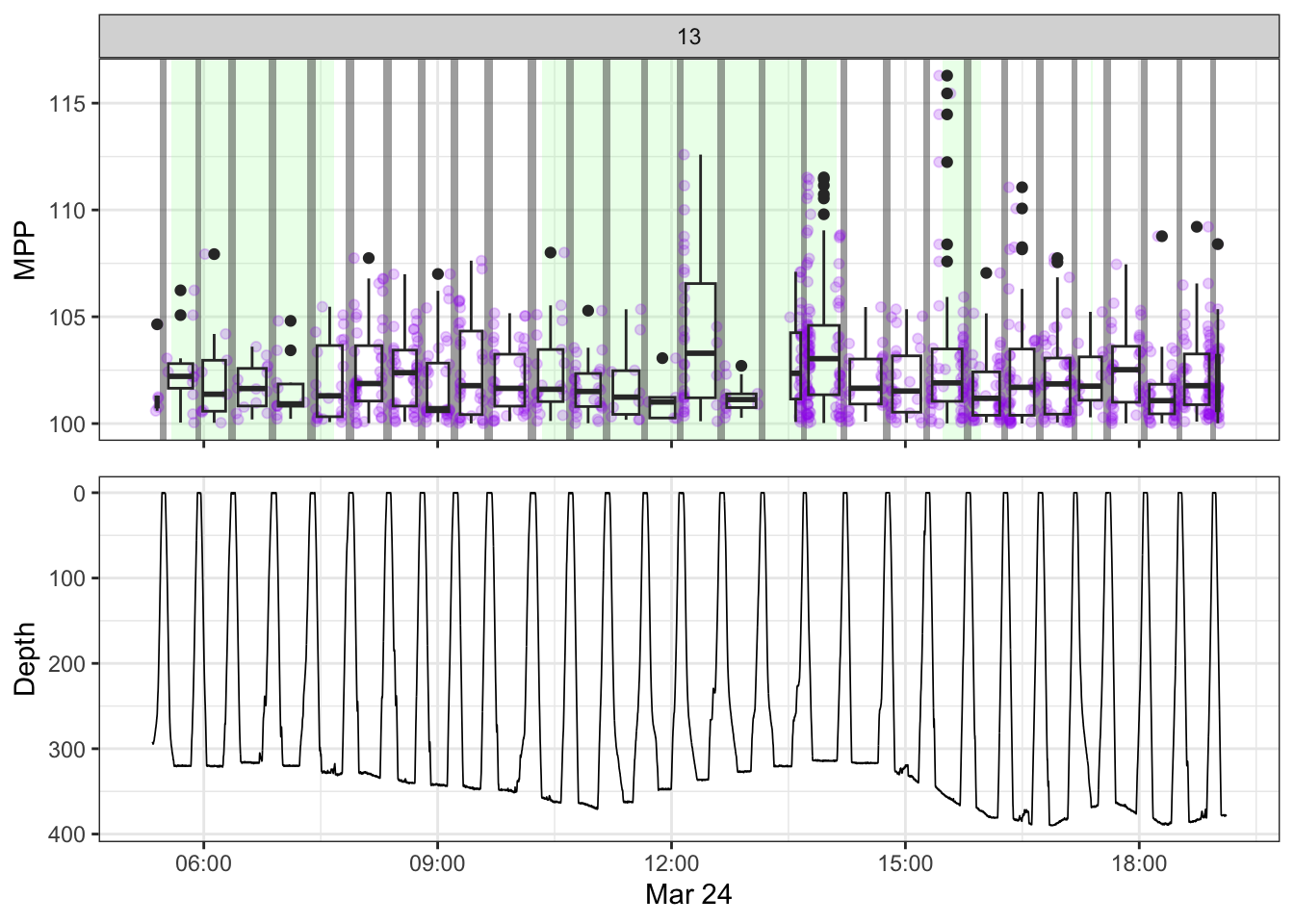
[[14]]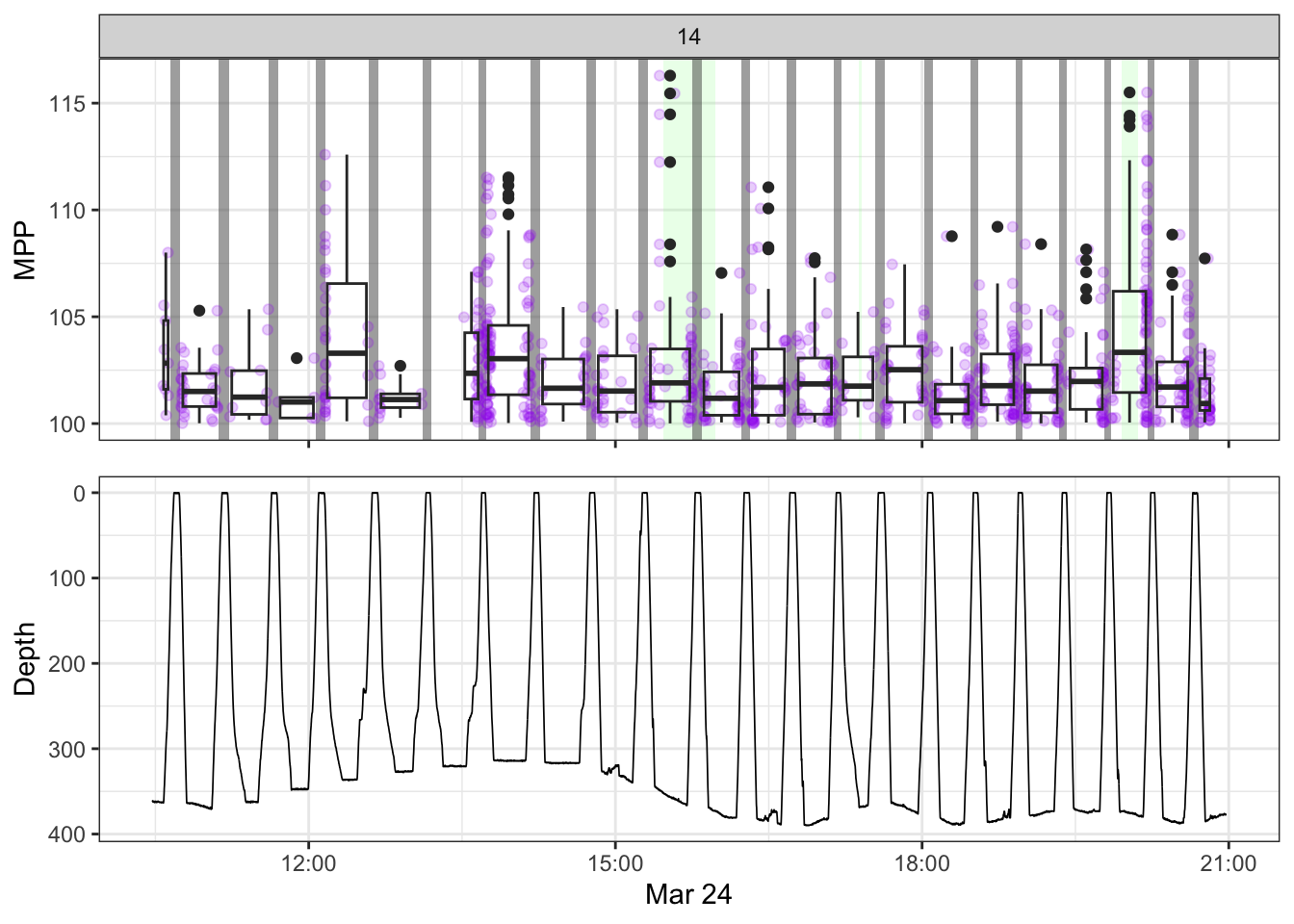
[[15]]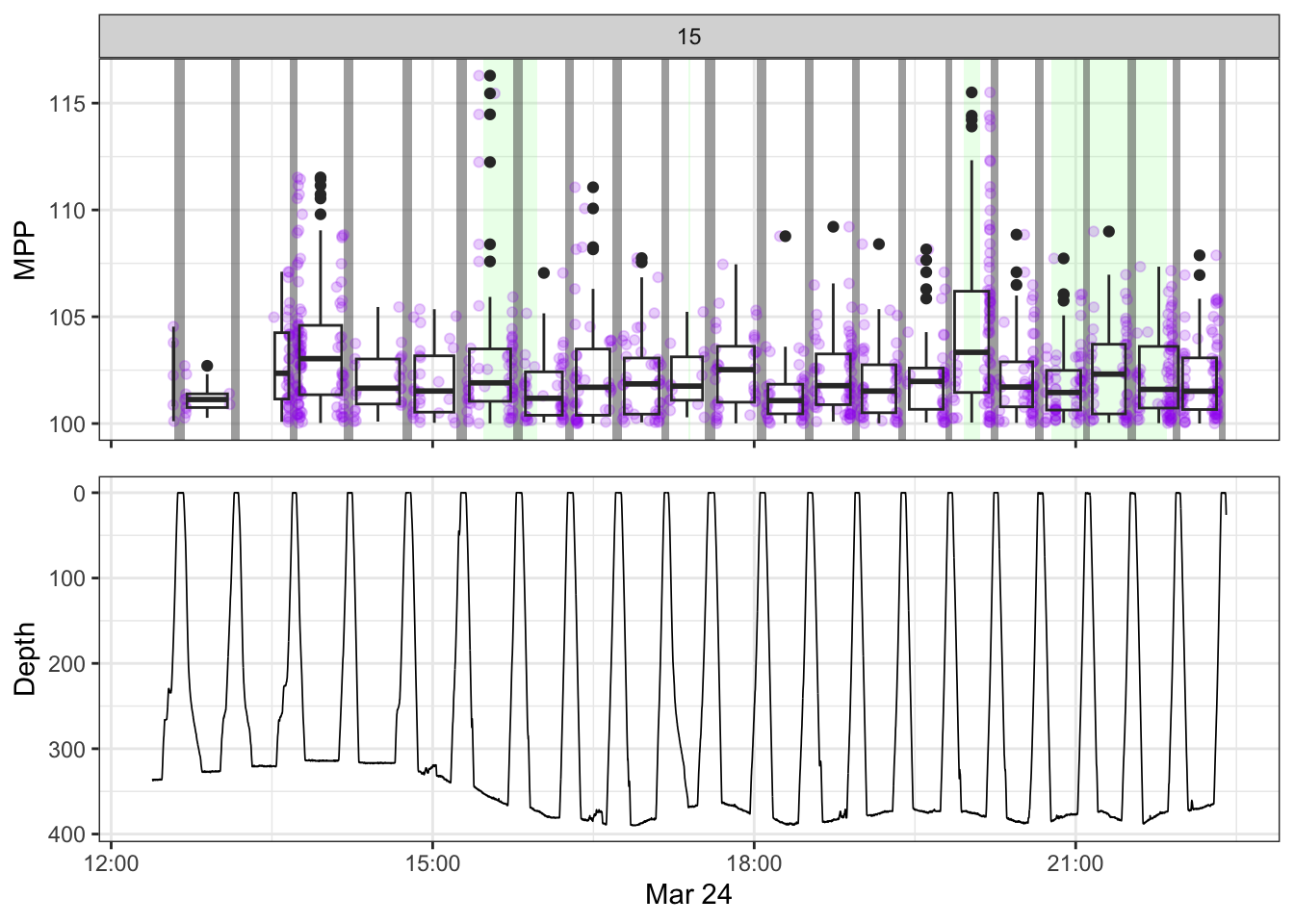
[[16]]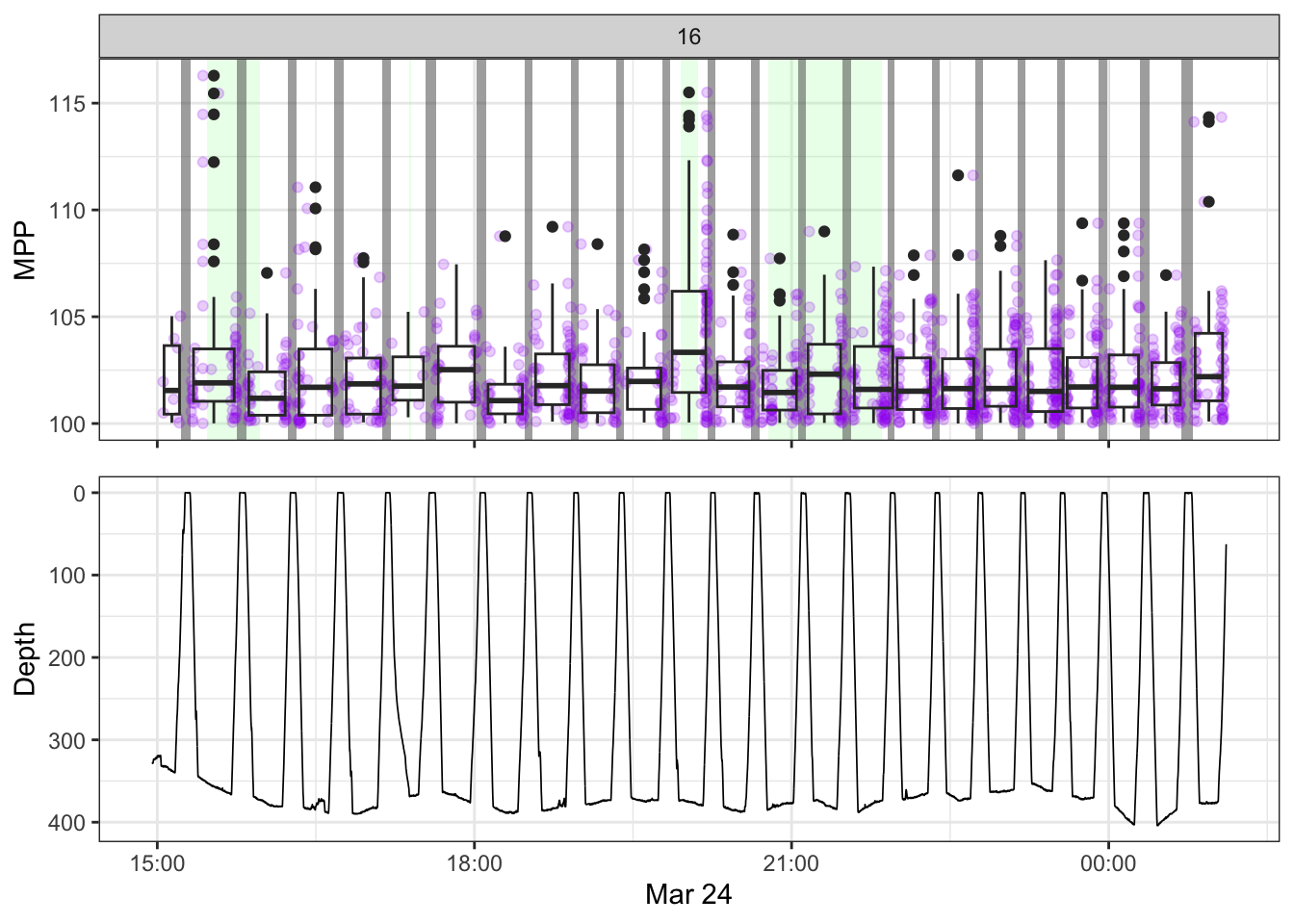
[[17]]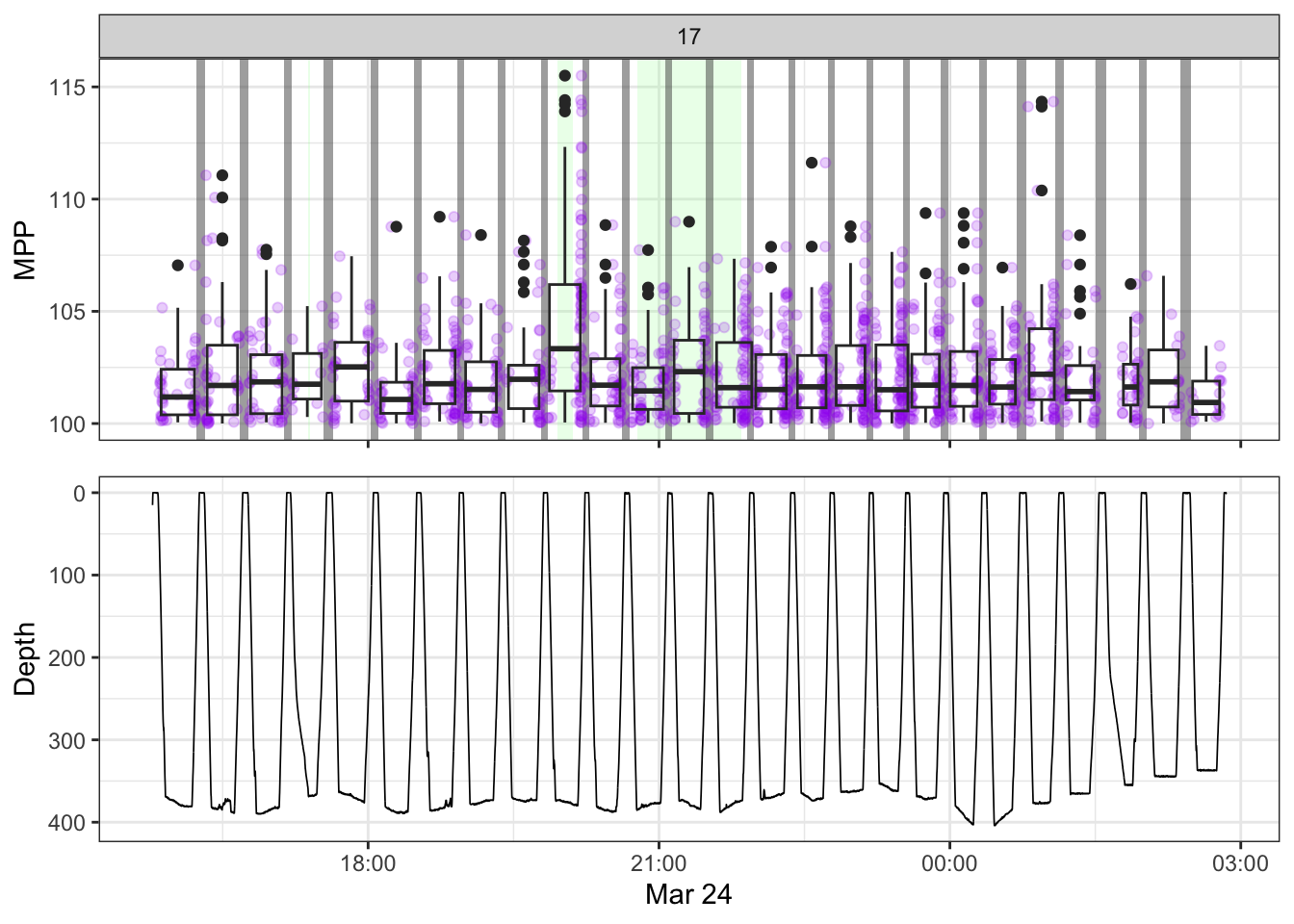
[[18]]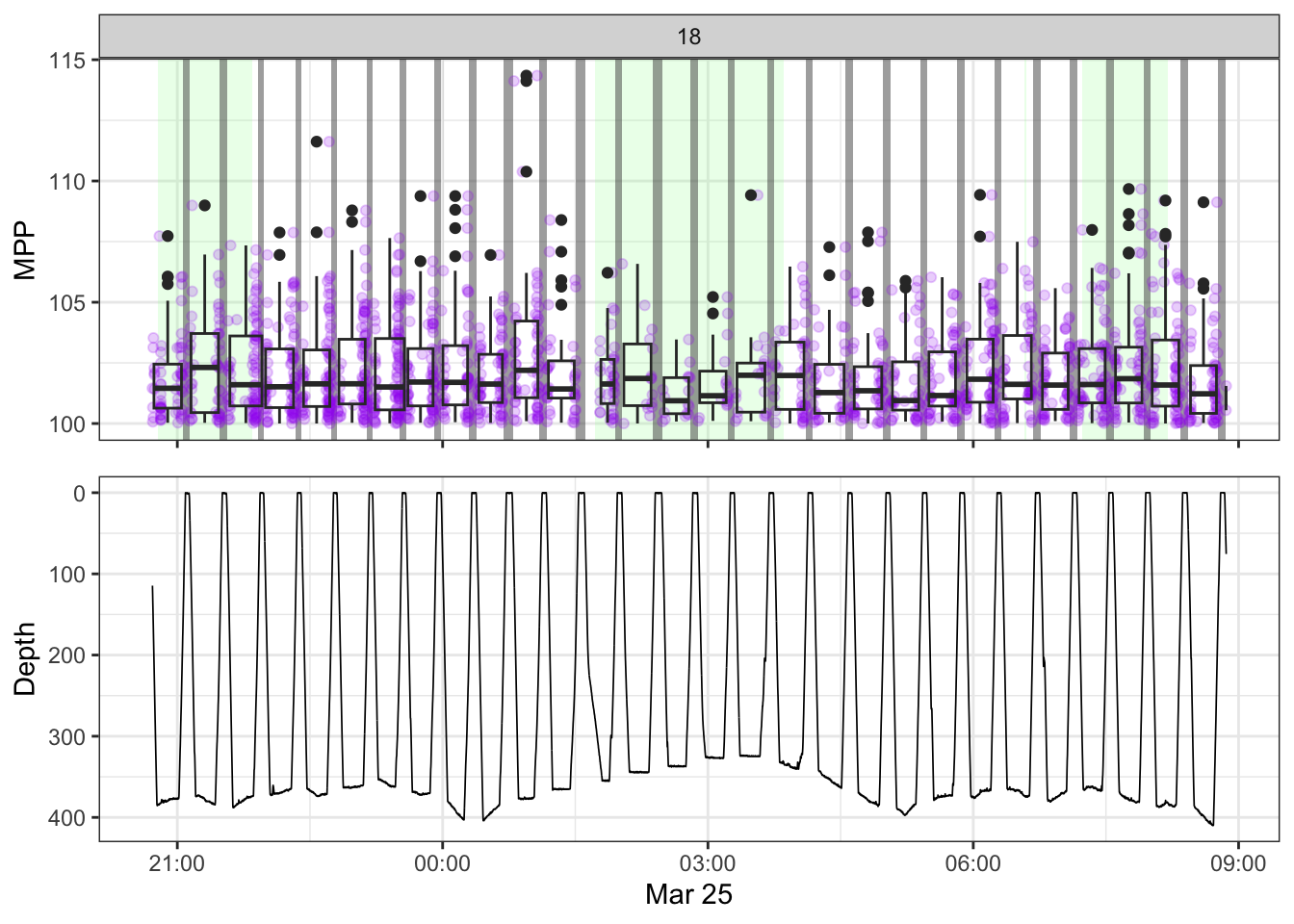
[[19]]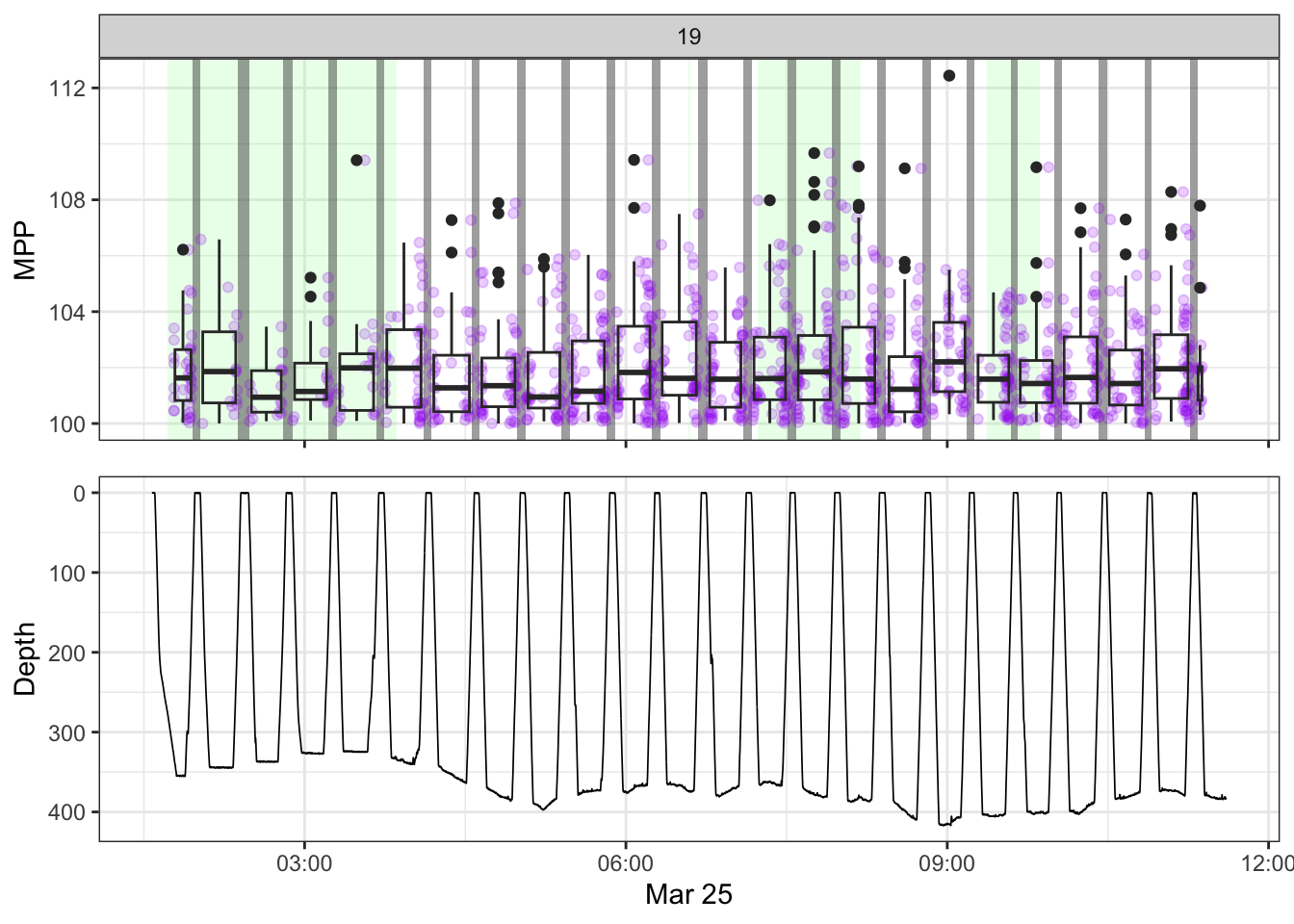
[[20]]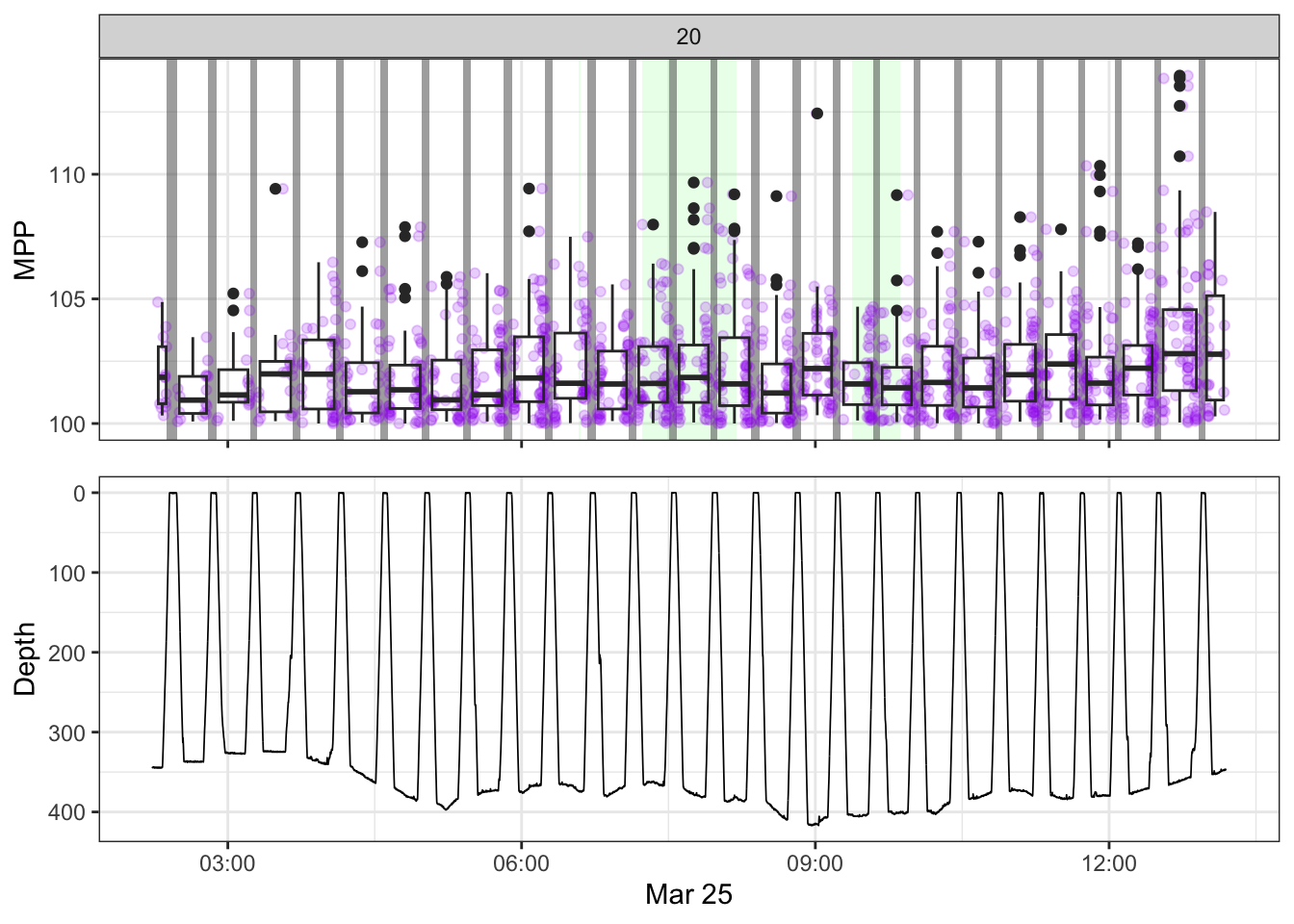
[[21]]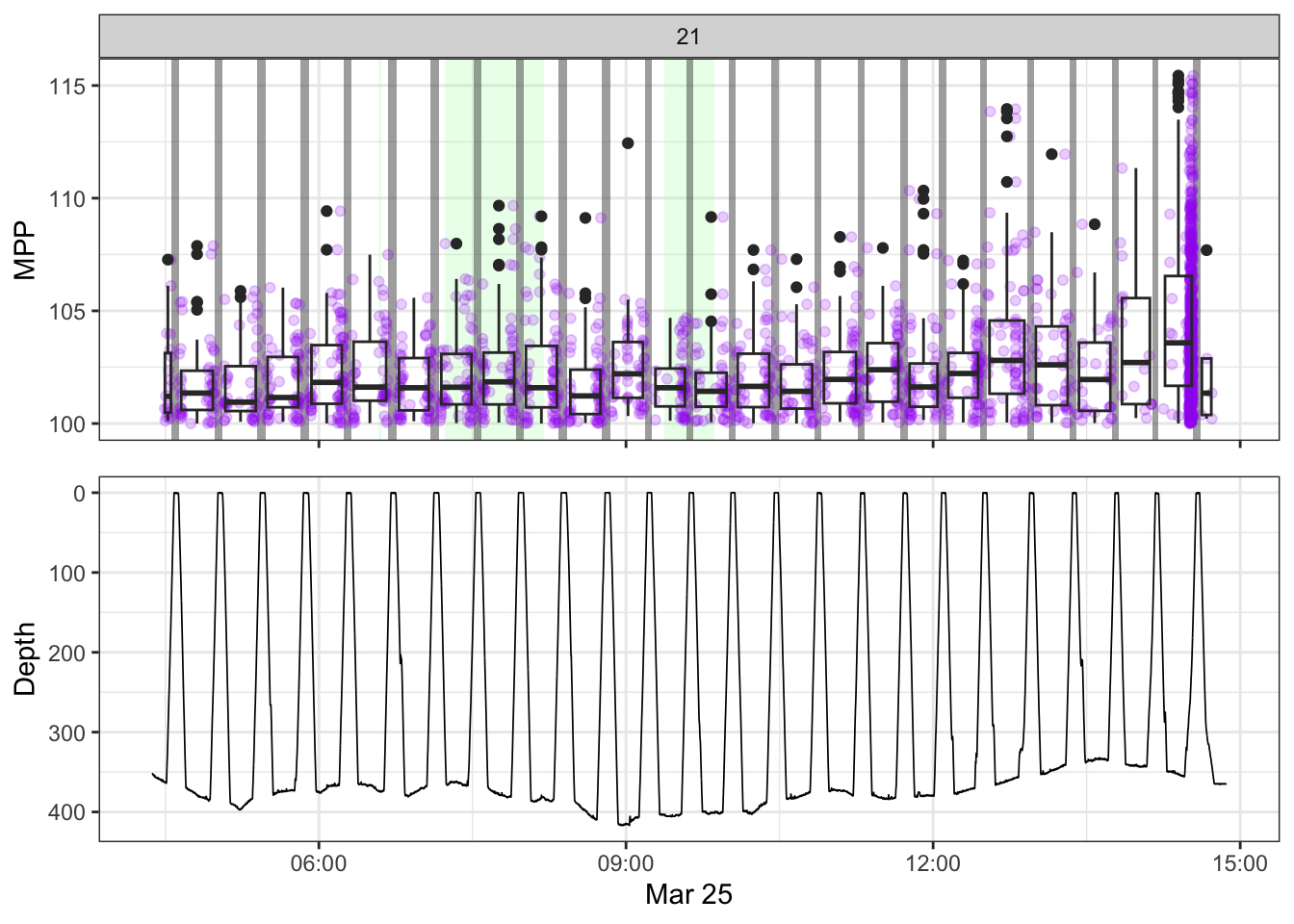
[[22]]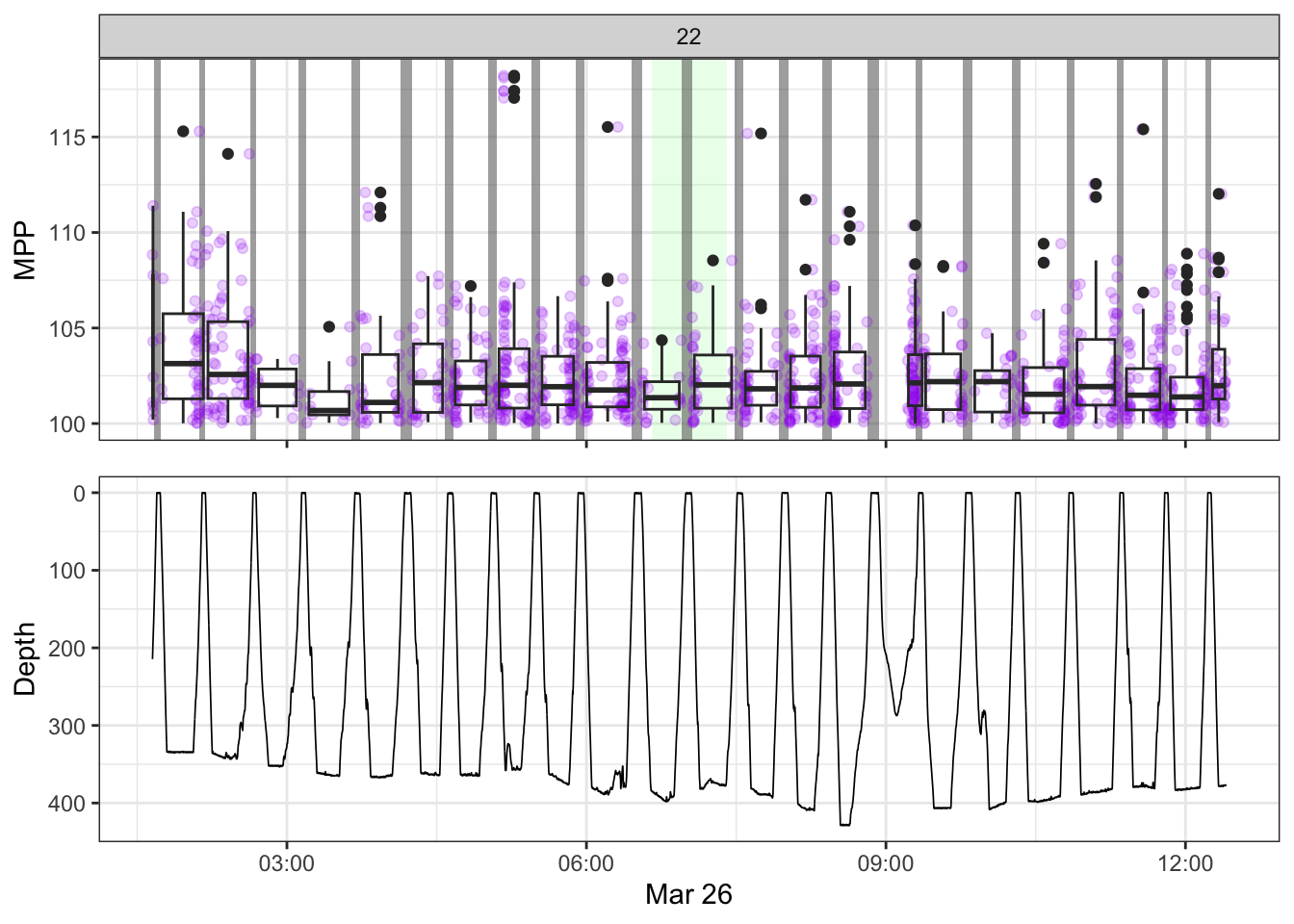
[[23]]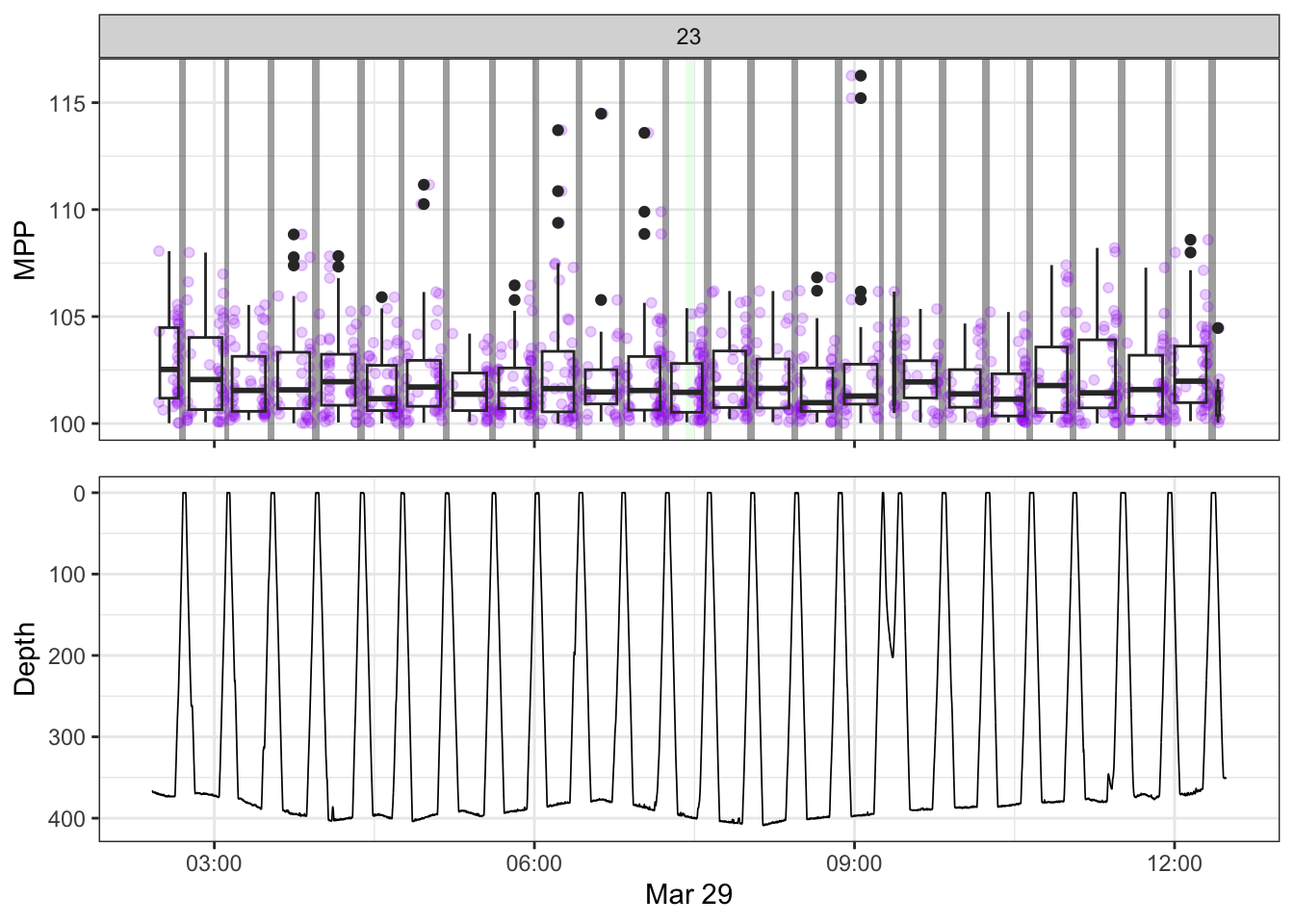
[[24]]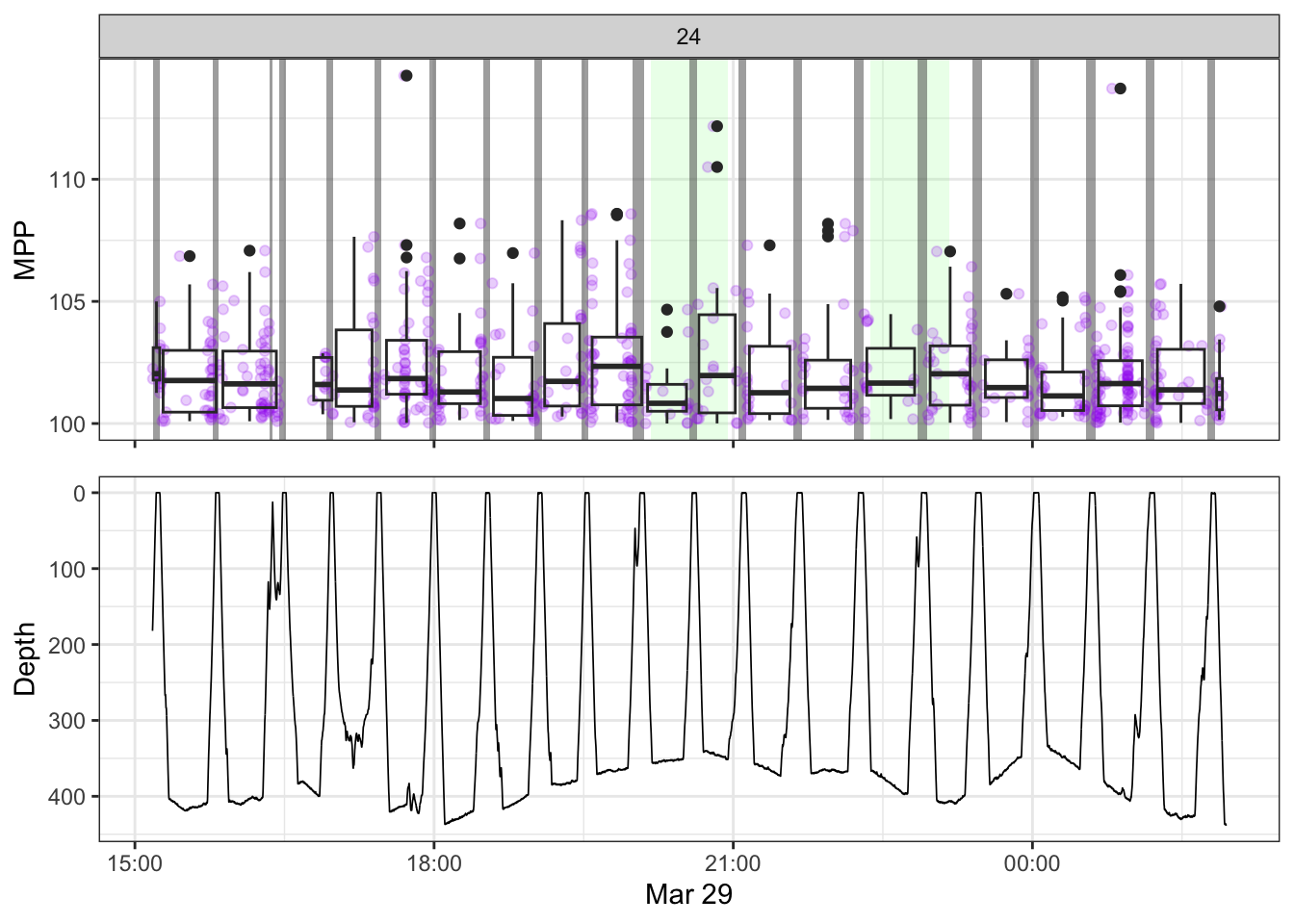
[[25]]
[[26]]
[[27]]
[[28]]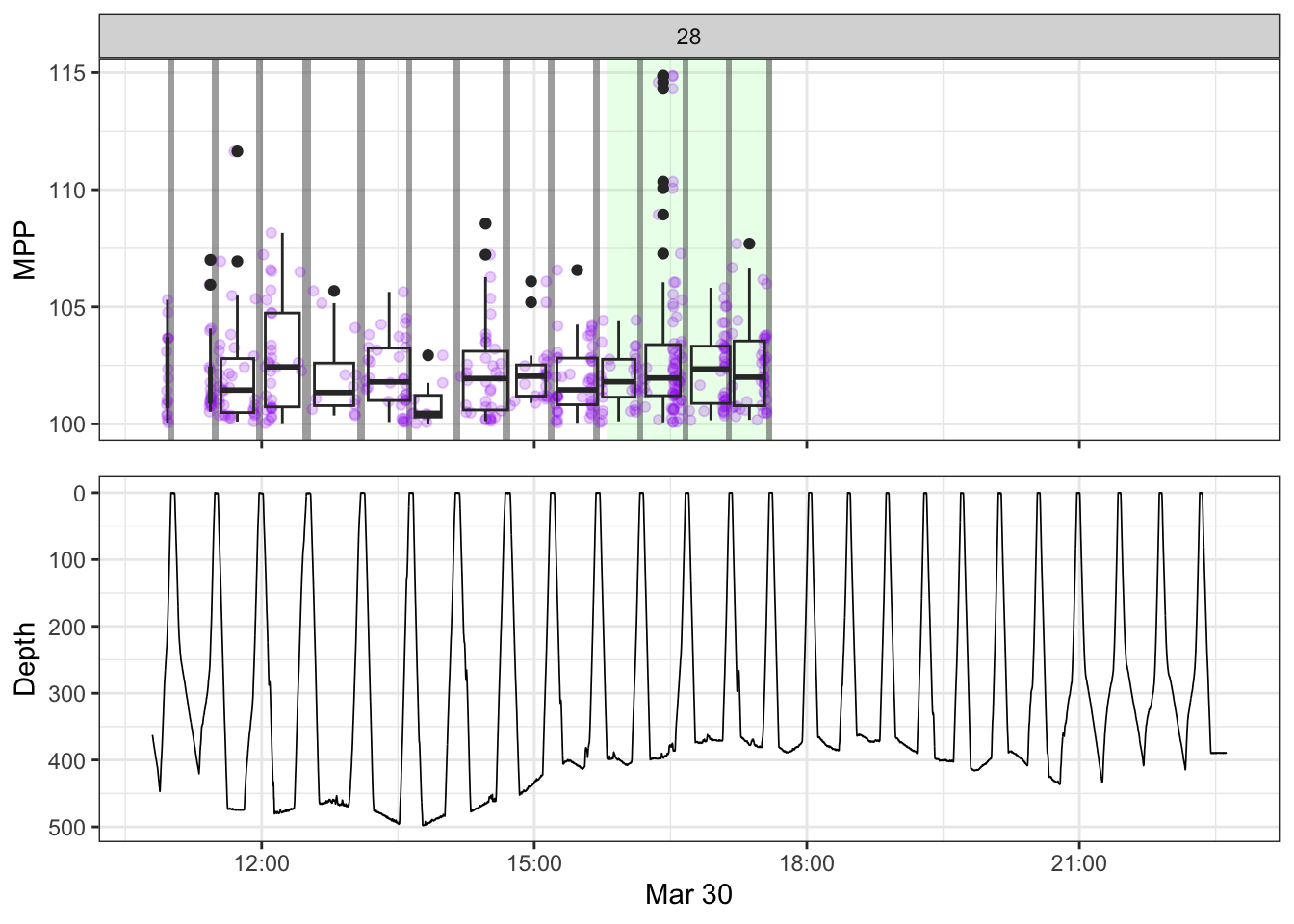
#save the plots to the outputs folder
# map(1:length(bout_plots),
# ~ggsave(paste0("output/bout_plots/comb_bouts/comb_bout_v3_", .x, ".png"),
# bout_plots[[.x]],
# width = 10,
# height = 6,
# units = "in"
# ))